No interaction is possible in batch mode. In batch execution mode, the pvbatch server runs an analysis script, written in Python, on the Blue Waters compute nodes. In interactive mode, the server component runs on the Blue Waters compute nodes and communicates with a client program running on a user's local machine. The server side of the application is a distributed memory MPI-based parallel application. ParaView supports client/server interactive mode as well as Python scripted batch mode. ParaView is a parallel visualization package based on the VTK C++ library. If this is only for you and you are already comfortable with using multiple applications and you don’t need to spend too much time viewing images and segmentations while looking at the CFD results then it may be simpler to use Paraview for final visualization.National Center for Supercomputing Applications at the University of Illinois at Urbana-Champaign Navigation 
What are your long term goals? Do you need tools that you use for your research or at some point you would want to have one integrated solution that others can use (potentially even clinicians)? If yes, then go all in on 3D Slicer is a good strategy. 
Slicer uses the same visualization toolkit as Paraview (VTK), so most visualization features are already available in Slicer, just not exposed on the user interface. You could either export images in a simplified format (resampled to have image axes aligned with patient axes, etc.) and live with Paraview’s limited image visualization features or import the CFD simulation results in to Slicer and add the visualization modes that you need in a small Python scripting module. Paraview is better for explorative visualizing of CFD results than Slicer, but only Slicer has all the tools that are required for your complete workflow (DICOM import, segmentation, rich image visualization, etc.). Is there some way I could access the data objects used for the visualization and save those instead? Just in general, is there a better way to export images and segmentations in the coordinate system of my choice?Īlso, all the data is aligned in the 3D render window. I can get around these issues, I just wonder if I’m making things harder than they have to be. Left side is contour from exported vtkOrientedImageData (hole outlined in blue), right side is contour from exported STL. Second, the vtkOrientedImageData seems to capture the bare minimum bounding box, which leads to holes not found in the STL: For one, the contour generated from it is of much poorer quality than the surface manually exported as an STL file. Is there a better way to retrieve the segmentation from Segment Editor (such as in a vtkPolyData format)? I could only retrieve a vtkOrientedImageData, and it has some issues. Is there a better way to retrieve the image orientations? This method seems to work, but has some problems: # use Contour Filter to extract surface in ParaView TransformF.SetInputData(orientedImageData) OrientedImageData = node.GetBinaryLabelmapInternalRepresentation('LevelSetSegmentationModel') Node = getNode('LevelSetSegmentationModel-segmentation') # retrieve the segmentation from Segment Editor node and transform it from RAS to IJK Transform_vtk.SetMatrix(ijk_to_lps_transform.flatten()) Image.GetIJKToRASDirections(orientations) Then I tried to use the Slicer Python Interpreter to transform the segmentation from RAS to IJK coordinates, using the API documentation: # get orientation and origin from the master volume I read this into ParaView and then removed the origin to have the image volume purely in IJK. nrrd format by default, which will be read into ParaView as a rectilinear grid in IJK coordinates. For now in ParaView, it seems easiest to use IJK coordinates. I want to export both of these into ParaView aligned for further postprocessing. 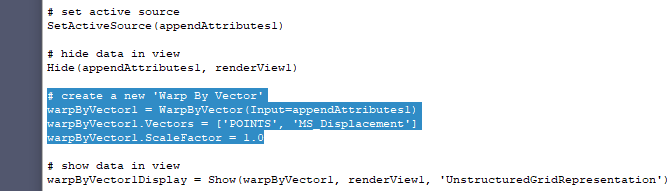
I have an image volume imported from a DICOM sequence which I’ve segmented first with the VMTK Extension (produces a node named “LevelSetSegmentationModel”) and then edited that segmentation with the Segment Editor (node “LevelSetSegmentationModel-segmentation”). I’m sure this type of problem pops up a lot, but I couldn’t find an exact answer to my problem. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
